# install.packages("motus",
# repos = c(birdscanada = 'https://birdscanada.r-universe.dev',
# CRAN = 'https://cloud.r-project.org'))
library(motus)
library(dplyr)
library(here)
library(DBI)
library(RSQLite)
library(forcats)
library(lubridate)
library(bioRad)
library(purrr)
library(ggplot2) Load & Format
Packages
You will need the motus package which you can download either online (Installing packages - Motus 2025) or from R.
Download
Note for authors:
You will have to separately run this R script that will follow the two next chunks before to continue. Indeed, `tagme( )` will prompt and require username and password that can’t be fed here.
Run this script to update database only. The procedure is heavy and takes a while to complete (up to 45 min).
# Global settings
setwd(dirname(rstudioapi::getSourceEditorContext()$path))
Sys.setenv(TZ="UTC")
proj.num <- 294
motusLogout()First, set general settings as working directory, Time Zone and Motus project number.
Then, make sure the environment is free from any connection to any other networked project to avoid undesired mistakes.
sql.motus <- tagme(projRecv = proj.num,
new = FALSE, # TRUE overwrites existing file
update = TRUE,
dir = here("qmd", "chapter_1","data"))
metadata(sql.motus, proj.num)
sql.motus <- dbConnect(SQLite(), here::here("qmd", "chapter_1", "data", "project-294.motus"))
df.alltags <- tbl(sql.motus, "alltags") %>%
dplyr::collect() %>%
as.data.frame() %>%
mutate(time = as_datetime(ts),
timeAus = as_datetime(ts, tz = "Australia/Sydney"),
dateAus = as_date(timeAus),
year = year(time),
doy = yday(time))Retrieve the data from Motus server to access them in a data-frame format.
motus::tagme()gets the data from the online Motus network, the ones part of your project (ie. ID = 294) only. Make sure you set your own directory. A file of type .sql is automatically created.
If you already have a .sql file, set new = FALSE and update = TRUE so you are updating your existing file instead of re-downloading the full thing, which can take a while.
motus::metadata() downloads the metadata from the online Motus network (receiver information and more).
motus::dbConnect() links your .sql to your environment. Avoid to use high memory as .sql is a lazy table and not yet hardly written on your hardware.
motus::tbl() extracts all the tags fat into your environment. Then formatted into a classic dataframe.
tail(df.alltags %>%
arrange(timeAus) %>%
select(timeAus, speciesEN, motusTagID, tagModel, pulseLen, recvDeployName, recv)) timeAus speciesEN motusTagID tagModel pulseLen
3452636 2026-02-27 10:32:27 <NA> 43288 NTQB2-6-2 2.5
3452637 2026-02-27 10:32:31 <NA> 43288 NTQB2-6-2 2.5
3452638 2026-02-27 10:32:34 <NA> 43288 NTQB2-6-2 2.5
3452639 2026-02-27 10:32:37 <NA> 43288 NTQB2-6-2 2.5
3452640 2026-02-27 14:29:47 Pied Stilt 60577 NTQB2-6-2 2.5
3452641 2026-02-27 14:31:05 Pied Stilt 60577 NTQB2-6-2 2.5
recvDeployName recv
3452636 Hexham Swamp SG-010DRPI3D708
3452637 Hexham Swamp SG-010DRPI3D708
3452638 Hexham Swamp SG-010DRPI3D708
3452639 Hexham Swamp SG-010DRPI3D708
3452640 Fullerton Entrance SG-A296RPI38A7E
3452641 Fullerton Entrance SG-A296RPI38A7EAbove, a quick view about the last data uploaded to Motus server, giving you the latest detection taken into account for this workflow.
Filter
SENSORGNOME
Receivers data and their meta-information must be filtered and formatted as well.
# Get summary
df.recvDeps <- tbl(sql.motus, "recvDeps") %>%
collect() %>%
as.data.frame() %>%
mutate(timeStart = as_datetime(tsStart),
timeStartAus = as_datetime(tsStart, tz = "Australia/Sydney"),
timeEnd = as_datetime(tsEnd),
timeEndAus = as_datetime(tsEnd, tz = "Australia/Sydney")) Indeed, some receivers might have been swap within the Motus array and along time, therefore the same SensorGnome ID might be used by multiple stations along time, leading to wrong results.
IMPORTANT: Make sure you tracked the history of your receivers to avoid misled results.
Here, we only need to rename the stations accurately and to remove those that are not used any longer.
# Correct the recv names (example)
station_rename <- list("Barry_Fullerton_cove" = "Fullerton Entrance",
"North Swann Pond" = "Swan Pond" ,
"Example_three" = "Example 3") # Apply corrections
df.recvDeps <- df.recvDeps %>%
mutate(name = recode(name, !!!station_rename)) %>%
rename(recvDeployName = "name")
# Filter not used stations as out of the local array
df.recvDeps <- df.recvDeps %>%
filter(!is.na(latitude),
recvDeployName != c("Throsby Creek Test Site"),
serno != c("SG-C621RPI3E17F", # test station
"SG-62A5RPI36710") ) # test station# Apply corrections
df.alltags <- df.alltags %>%
mutate(recvDeployName = recode(recvDeployName, !!!station_rename))Survey time frame
df.recvDeps <- df.recvDeps %>% filter(timeStartAus > "2023-01-31 00:00:00 AEDT")Finally, we need to set temporal aspect of the survey effort.
Day one for our local Motus array to start listening any tag signals is set on the 31st January 2023 - one month before the first day a bird has been caught and tagged (information found into the SharePoint to access the file.).
Import band ID
As we might re-trap individuals and re-tag them with a new MOTUS tag, we can’t use the Motus tag ID as unique ID anymore and have to use Band ID which supposes to last on the same bird from banding date to individual’s death. Unfortunately, the Band ID is not imported by MOTUS network at the moment. So, we need to import it from our own record accessible on SharePoint here, download the .csv.
# Load df with date at the beginning
spreadsheet <- read.csv(here::here( "qmd", "chapter_1", "data", "spreadsheet", paste0(Sys.Date(), "-teams.sheet.csv"))) %>%
# Keep only the tagged ones
filter(Radio.tag. == "Y") %>%
# Variable names
rename(DateAUS.Trap = "Date",
motusTagID = "Motus.tag.ID",
speciesEN = "Species") %>%
# Value names
mutate(speciesEN = case_when(
speciesEN == "Eastern Curlew" ~ "Far Eastern Curlew",
speciesEN == "Black-winged Stilt" ~ "Pied Stilt",
speciesEN == "Pacific Golden Plover" ~ "Pacific Golden-Plover",
speciesEN == "Whimbrel" ~ "Eurasian Whimbrel",
TRUE ~ speciesEN )) %>%
# Format
mutate(motusTagID = as.factor(motusTagID),
DateAUS.Trap = as.Date(DateAUS.Trap),
Band.ID = as.factor(Band.ID)) %>%
select(Band.ID, motusTagID, speciesEN, DateAUS.Trap, everything())
# Format motusTagID for further merging
df.alltags <- df.alltags %>%
mutate(motusTagID = as.factor(motusTagID))
# Join unique Band IDs for inconsistent motusTag (same bird re-tagged, etc)
df.alltags <- left_join(df.alltags,
spreadsheet %>%
filter(is.na(Euthanised.)) %>%
select(motusTagID, DateAUS.Trap, Band.ID, Bander),
by = "motusTagID")So we merged our records with Band ID for each bird to the Motus data, linked with the Motus tag ID. Allowing unique individual IDs.
Add key variables
It is crucial for our further analysis to add the Tidal and Circadian cycles as variables so we can match each bird detection with a categorical value for a date, a tide (high/low) and a period of the day (night/day).
To do so, we must first fetch the tide data.
# Read tide.csv
tideData <- read.csv(here("qmd", "chapter_1", "data", "tides", "TideDataNewcastle.csv"))
# Format date and datetime columns
tideData <- tideData %>% mutate(
date = dmy(date, tz = "Australia/Sydney"),
tideDateTimeAus = dmy_hm(tideDateTimeAus, tz = "Australia/Sydney")
)
# Classify tides as diurnal or nocturnal
tideData <- tideData %>% mutate(
sunriseNewc = sunrise(date, 151.7833, -32.9167, elev = -0.268, tz = "Australia/Sydney", force_tz = TRUE),
sunsetNewc = sunset(date, 151.7833, -32.9167, elev = -0.268, tz = "Australia/Sydney", force_tz = TRUE),
sunriseNewcTime = strftime(sunriseNewc, format = "%H:%M:%S", tz = "Australia/Sydney"),
sunsetNewcTime = strftime(sunsetNewc, format = "%H:%M:%S", tz = "Australia/Sydney")
)
# Define either diurnal or nocturnal
tideData <- tideData %>% mutate(
day_night = case_when(
tideDateTimeAus >= sunriseNewc & tideDateTimeAus <= sunsetNewc ~ "Diurnal",
TRUE ~ "Nocturnal"
)
)
# Categorise each tide by tidal/diel period
tideData <- tideData %>% mutate(
tideCategory = case_when(
high_low == "Low" & day_night == "Diurnal" ~ "Diurnal_Low",
high_low == "Low" & day_night == "Nocturnal" ~ "Nocturnal_Low",
high_low == "High" & day_night == "Diurnal" ~ "Diurnal_High",
high_low == "High" & day_night == "Nocturnal" ~ "Nocturnal_High") %>%
as_factor())
# Add numeric ID to each category, allowing for unique tide bins
tideData <- tideData %>%
group_by(tideCategory) %>%
mutate(tideID = paste0(tideCategory, "_", row_number())) %>%
ungroup()Note that tide data-set must be extracted from New South Wales government resources and formatted into a .csv file - made for you and directly accessible here.
All the Tide categories are set for a unique point in Newcastle, easing the complexity of small scale differences for the tide within the estuary.
Let’s now add our key variables for each bird detection:
Signal strength
Circadian period
Tidal period
# Load useful functions from Callum Gapes work
tidalCurve <- readRDS(here::here("qmd", "chapter_1", "data", "tides", "tidalCurve.rds"))
tidalCurveFunc <- splinefun(tideData$tideDateTimeAus, tideData$tideHeight, method = "natural")
get.tideIndex <- function(time){ return(which.min(abs(tideData$tideDateTimeAus-time)))}
# Add key variables
df.alltags <- df.alltags %>%
# Positive signal strength (min. = 0) for plotting
mutate(sigPositive = sig + abs(min(sig))) %>%
# Sunrise/set
sunRiseSet(lat = "recvDeployLat",
lon = "recvDeployLon",
ts = "ts") %>%
mutate(sunriseNewc = sunrise(dateAus, 151.7833, -32.9167, elev = -0.268, tz = "Australia/Sydney", force_tz = TRUE),
sunsetNewc = sunset(dateAus, 151.7833, -32.9167, elev = -0.268, tz = "Australia/Sydney", force_tz = TRUE)) %>%
# Tide
mutate(tideHeight = tidalCurveFunc(timeAus),
tideIndex = map_dbl(timeAus, get.tideIndex))
tide_values <- tideData[df.alltags$tideIndex,
c("tideDateTimeAus",
"high_low",
"day_night",
"tideCategory",
"tideID",
"tideHeight")]
# Stick and factorise the variables
df.alltags <- df.alltags %>%
mutate(tideDateTimeAus = tide_values$tideDateTimeAus,
tideHighLow = as_factor(tide_values$high_low),
tideDiel = as_factor(tide_values$day_night),
tideCategory = as_factor(tide_values$tideCategory),
tideCategoryHeight = tide_values$tideHeight,
tideID = as_factor(tide_values$tideID),
tideTimeDiff = abs(difftime(timeAus, tideDateTimeAus, units = "hours")))
df.alltags <- df.alltags %>%
mutate(Band.ID = as.factor(Band.ID))Save
Note for authors:
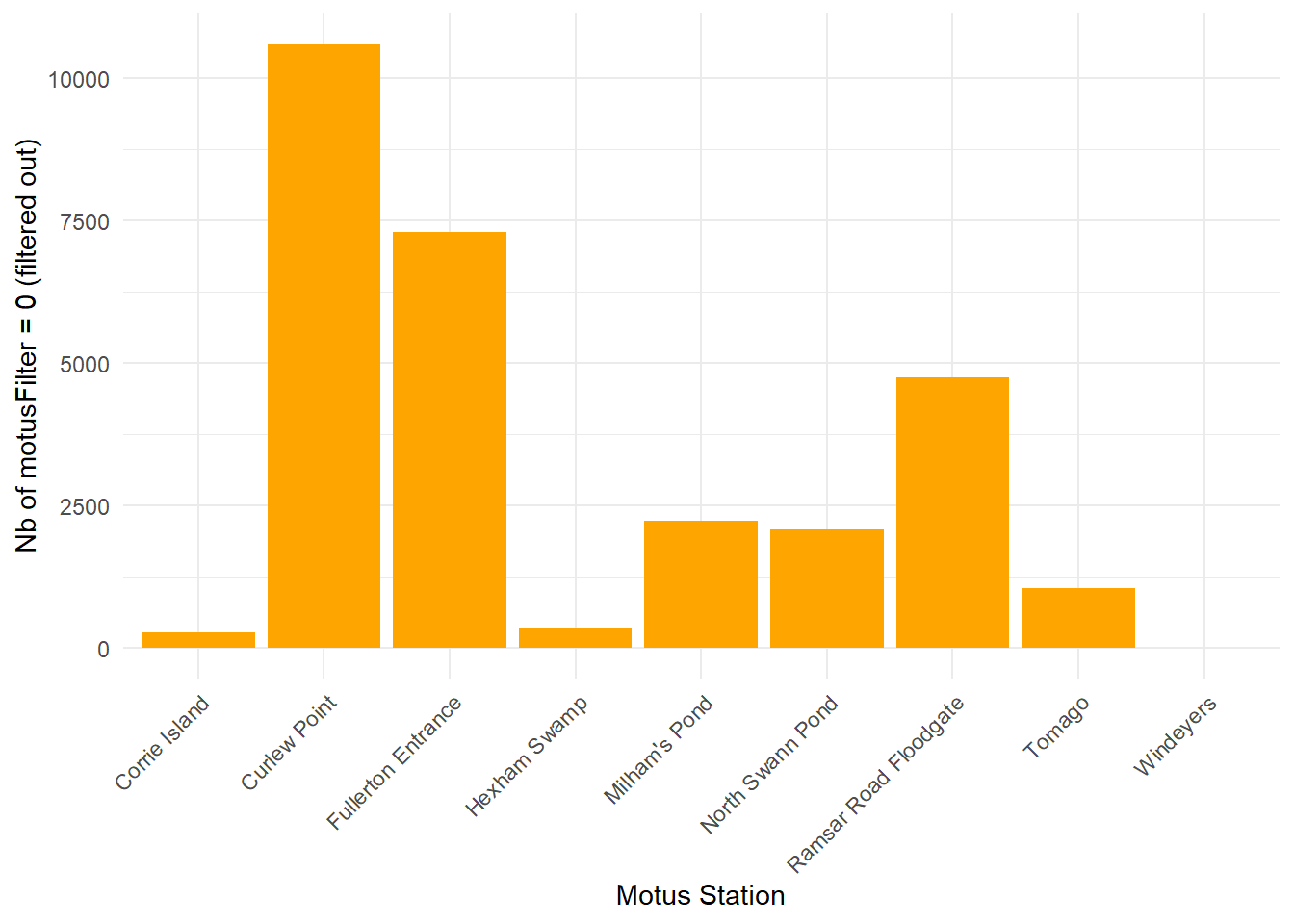
Some observations (n = 11130) in the detections data retrieved from Motus server do hold NA values for speciesEN.
This can occur when tags are activated (regardless whether they are deployed on Motus server) during a catch event within the vicinity of a deployed and operational Motus station.
Catching at Curlew point on 10 March 2026 generated about 11 100 tag detections from tags we did not deployed.
These are removed from the data-set.
# Bird detection dqtq
saveRDS(df.alltags, here::here("qmd", "chapter_1", "data", "motus", paste0(Sys.Date(), "-data", ".rds" )))
# Receiver information
saveRDS(df.recvDeps, here::here("qmd", "chapter_1", "data", "motus", paste0(Sys.Date(), "-recv-info", ".rds" )))
# Spreadsheet tracking BandID
saveRDS(spreadsheet, here::here("qmd", "chapter_1", "data", "spreadsheet", paste0(Sys.Date(), "-spreadsheet", ".rds" )))
# Tide tables
saveRDS(tideData, here("qmd", "chapter_1", "data", "tides", "tideData.rds"))Save your formatted data, now ready for analysis!
Reproducibility
Ready-to-Go!
To reproduce any of the analysis of this Research Project, simply click and download the data (a password might be required - maxime.marini@uon.edu.au).
Citation: Griffin, A. Shorebird monitoring in central NSW estuaries (Project 294). 2019. Data accessed from Motus Wildlife Tracking System, Birds Canada. Available: https://motus.org/. Accessed: YYYY-MM-DD
Once loaded in your R environment, the file will display five objects:
sql.motus: raw motus file from which you can call multiple tables (see documentation)data_all: contains all the raw detections for each Lotek nanotagsrecv: contains all the details for each receiver of the Motus arrayspreadsheet: contains all the capture events details provided for each trapped individualtideData: tidal details for Newcaslte, NSW
The data are last updated on the 21 December 2025.