library(dplyr)
library(here)
library(ggplot2)
library(lubridate)
library(tidyr)Estuary usage - days
Variables
Load your data in your R environment - see Load & Format > Reproducibility
Let’s count days and detecting stations for each bird.
# Get deployment duration for each tags (in days)
tag_dep_duration <- data_all %>%
group_by(Band.ID) %>%
summarise(start_date = min(DateAUS.Trap),
end_date = max(dateAus),
period_tag_dep_d = 1 + time_length(interval(start = start_date, end = end_date), unit = "day"), # +1 to include the trapping day
.groups = "drop") %>%
select(Band.ID, period_tag_dep_d)
# Get number days each tag has been detected at least once in any station
tag_dep_nb_d <- data_all %>%
mutate(dateAus = as.Date(dateAus)) %>%
group_by(Band.ID) %>%
summarise(days_detect = n_distinct(dateAus), .groups = "drop")
# Get number of days one tag has been recorded in >1 station (for each tags)
multiple_site_detect <- data_all %>%
group_by(Band.ID, dateAus) %>%
# Counts per day the number of station one tag has been recorded
summarise(distinct_sites = n_distinct(recvDeployName),
.groups = "drop_last") %>%
# Tells if one day one tag has been recorded several sites
mutate(multiple_detect = distinct_sites > 1) %>%
group_by(Band.ID) %>%
summarise(multiple_site_detect = sum(multiple_detect),
.groups = "drop")We generate tag_dep_duration that counts the total of days period_tag_dep_d each bird Band.ID had a tag on and activated. We make tag_dep_nb_d that quantifies the number of days days_detect each bird have been detected at least once in the Motus array.
So, we can then compare the first value with the second, and obtain our metric of interest in percentage.
Additionally, we quantified in multiple_site_detect the number of days each bird has been detected in multiple stations.
Results
INDIVIDUALS
# Get number of day each tag has been recorded /species and /station
tag_detection_indiv <- data_all %>%
group_by(Band.ID, recvDeployName, speciesEN) %>%
# to count sp*ID per recv
reframe(days_recorded = n_distinct(dateAus)) %>%
# flip table to get one column per station
pivot_wider(names_from = recvDeployName,
values_from = days_recorded,
values_fill = 0) %>% # = 0 if NA
left_join(multiple_site_detect, by = "Band.ID") %>%
left_join(tag_dep_nb_d, by = "Band.ID") %>%
left_join(tag_dep_duration, by = "Band.ID") %>%
mutate(perc = round((days_detect / period_tag_dep_d) * 100, 1)) # nb of day one tag is recorded once / nb of day the tag is deployedThe above table presents daily detection for each bird and for each Motus station in the local array:
multiple_site_detect: number of days a bird has been detected in more than one stationdays_detect: number of days a bird has been detected in at least one stationperiod_tag_dep_d: total of days a bird was detectable with a tag on and activatedperc: (days_detect/period_tag_dep_d) x 100
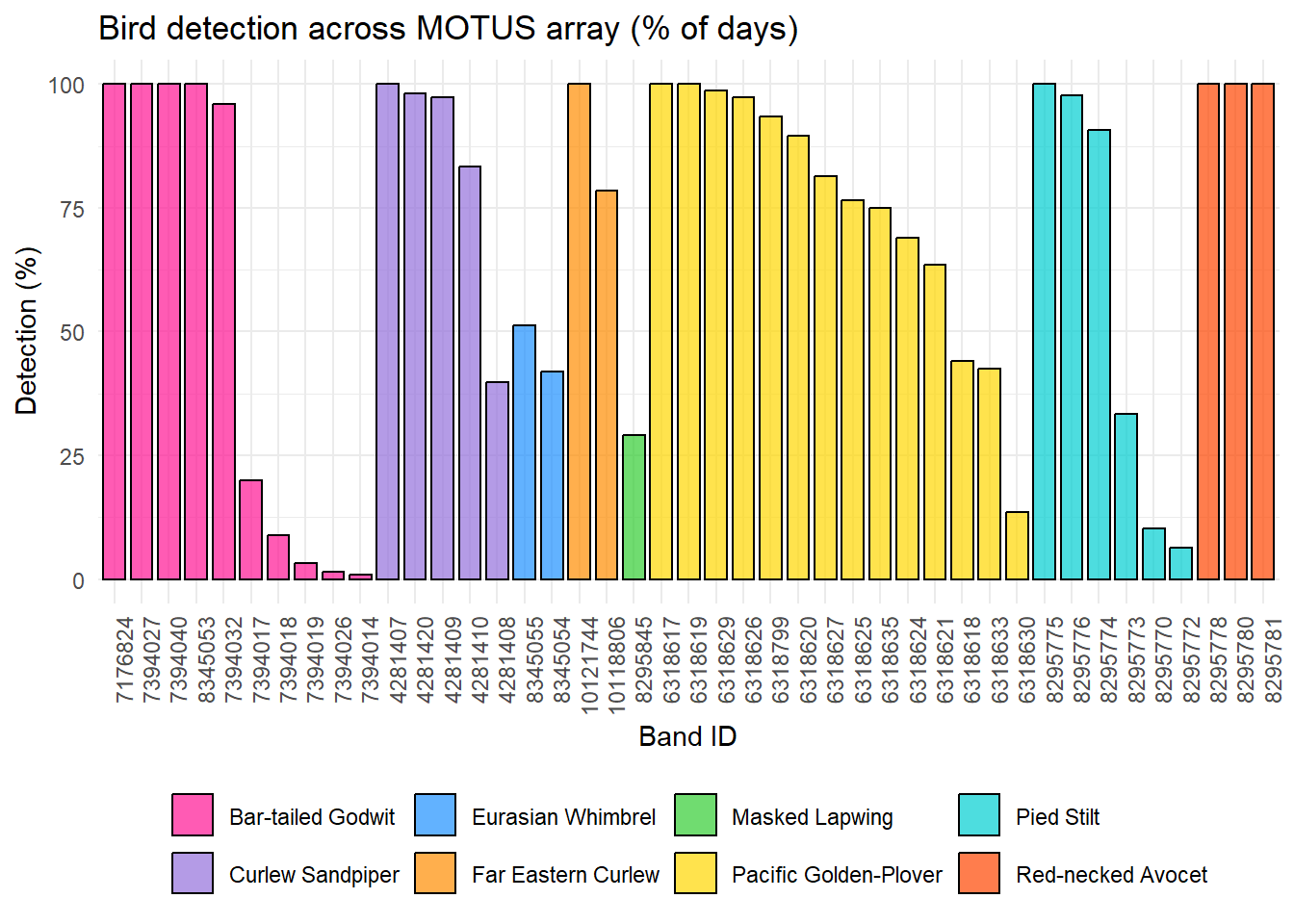
Visualise below for each birds the number of days the individual has been detected at least once and in at least one Motus station in the array, against the total of days the bird had a tag on and activated (results in %).
SPECIES
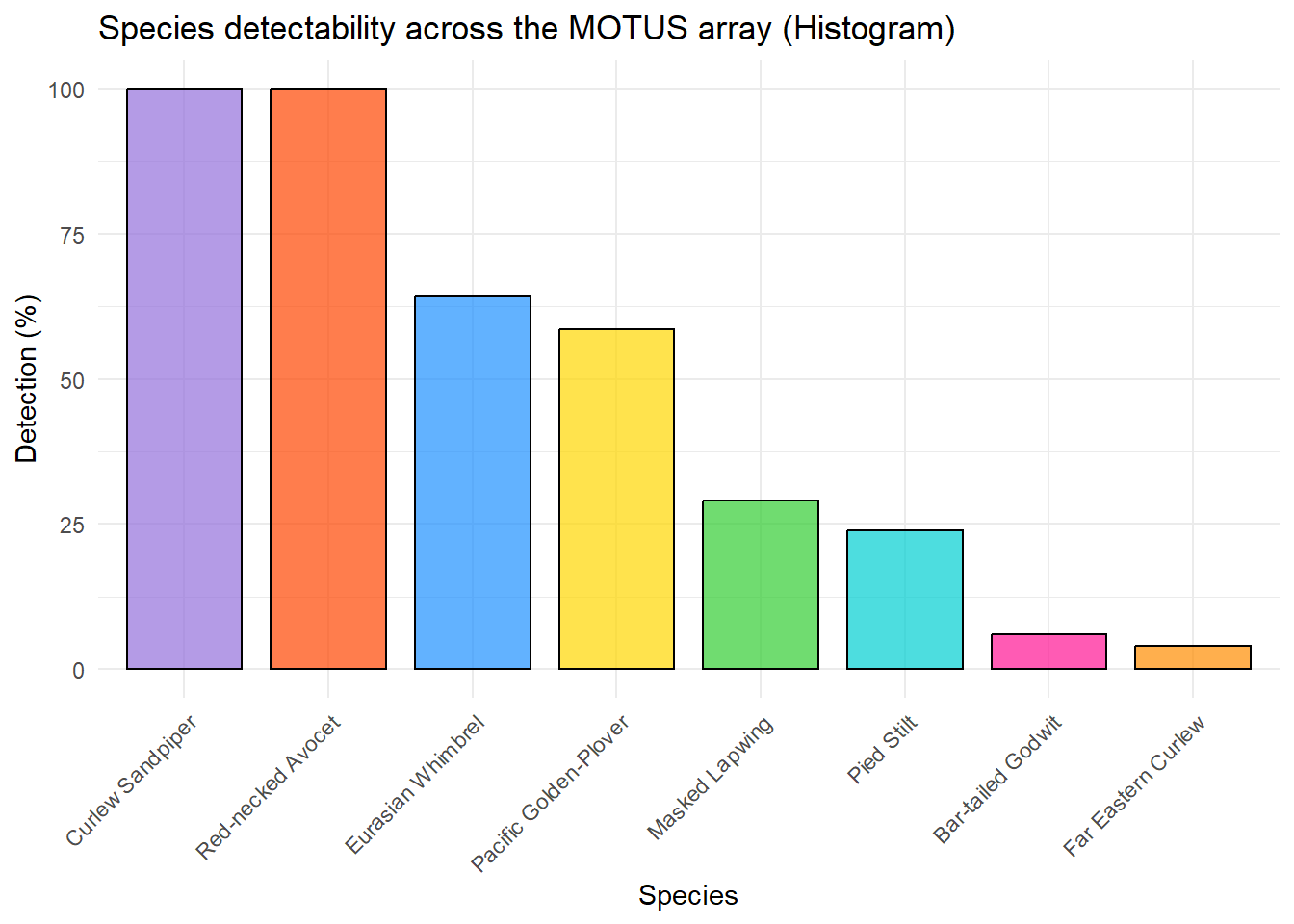
Below are the same results as above, but for each species. So the data has been grouped by species.
| speciesEN | Curlew Point | Fullerton Entrance | Hexham Swamp | Milhams Pond | Ramsar Road | Swan Pond | Tomago | Windeyers | multiple_site_detect | days_detect | period_tag_dep_d | perc |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Bar-tailed Godwit | 48 | 50 | 0 | 5 | 0 | 0 | 22 | 0 | 43 | 64 | 1064 | 6.0 |
| Curlew Sandpiper | 106 | 52 | 0 | 7 | 62 | 107 | 65 | 0 | 107 | 111 | 111 | 100.0 |
| Eurasian Whimbrel | 36 | 22 | 0 | 0 | 0 | 0 | 9 | 0 | 16 | 50 | 78 | 64.1 |
| Far Eastern Curlew | 1 | 25 | 0 | 22 | 0 | 0 | 12 | 0 | 17 | 43 | 1078 | 4.0 |
| Masked Lapwing | 9 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 9 | 31 | 29.0 |
| Pacific Golden-Plover | 219 | 77 | 51 | 77 | 76 | 96 | 1 | 17 | 158 | 280 | 478 | 58.6 |
| Pied Stilt | 18 | 105 | 0 | 2 | 69 | 61 | 0 | 0 | 68 | 159 | 664 | 23.9 |
| Red-necked Avocet | 24 | 29 | 3 | 4 | 0 | 5 | 21 | 0 | 29 | 29 | 29 | 100.0 |
ggplot(tag_detection_indiv %>%
add_count(speciesEN) %>%
mutate(species_label = paste0(speciesEN, " (n = ", n, ")")),
aes(x = reorder(species_label, -perc, FUN = median),
y = perc, fill = speciesEN)) +
geom_boxplot(width = 0.8, alpha = 0.7, color = "black") +
scale_fill_manual(values = species_colors) +
theme_minimal() +
theme(axis.text.x = element_text(angle = 45, hjust = 1),
legend.position = "none") +
labs(x = "Species (n = number of individuals)",
y = "Detection (%)",
title = "Species detectability across the MOTUS array (Boxplot)")
detection_table <- tag_detection_indiv %>%
add_count(speciesEN) %>%
mutate(Species = speciesEN) %>%
group_by(Species, speciesEN) %>%
summarise(
n_individuals = n(),
min = round(min(perc, na.rm = TRUE), 1),
q1 = round(quantile(perc, 0.25, na.rm = TRUE),1),
median = round(median(perc, na.rm = TRUE),1),
q3 = round(quantile(perc, 0.75, na.rm = TRUE),1),
max = round(max(perc, na.rm = TRUE),1),
mean = round(mean(perc, na.rm = TRUE),1),
sd = round(sd(perc, na.rm = TRUE),1),
.groups = 'drop'
) %>%
arrange(desc(median)) %>%
select(-speciesEN)
library(gt)
detection_table %>%
gt() %>%
opt_align_table_header(align = "left") %>%
tab_style(
style = cell_text(weight = "bold"),
locations = cells_column_labels() ) %>%
tab_style(
style = cell_text(style = "italic"),
locations = cells_body(columns = c(Species))) | Species | n_individuals | min | q1 | median | q3 | max | mean | sd |
|---|---|---|---|---|---|---|---|---|
| Red-necked Avocet | 3 | 100.0 | 100.0 | 100.0 | 100.0 | 100.0 | 100.0 | 0.0 |
| Curlew Sandpiper | 5 | 39.7 | 83.3 | 97.3 | 98.2 | 100.0 | 83.7 | 25.5 |
| Far Eastern Curlew | 2 | 78.4 | 83.8 | 89.2 | 94.6 | 100.0 | 89.2 | 15.3 |
| Pacific Golden-Plover | 14 | 13.6 | 65.0 | 79.0 | 96.4 | 100.0 | 74.7 | 26.1 |
| Pied Stilt | 6 | 6.3 | 16.0 | 62.0 | 96.0 | 100.0 | 56.4 | 44.7 |
| Bar-tailed Godwit | 10 | 0.8 | 4.7 | 58.0 | 100.0 | 100.0 | 53.0 | 49.0 |
| Eurasian Whimbrel | 2 | 41.9 | 44.2 | 46.6 | 48.9 | 51.3 | 46.6 | 6.6 |
| Masked Lapwing | 1 | 29.0 | 29.0 | 29.0 | 29.0 | 29.0 | 29.0 | NA |